Purpose
This project is part of ACDN and enables multiple hospitals to collaboratively develop machine learning models while maintaining patient privacy. It uses Federated Learning (FL): each hospital trains locally on its de-identified datasets, while only numeric model updates are exchanged with a central server for aggregation.
No Patient Health Information (PHI) is transferred to the FL server. Raw clinical datasets remain within each hospital.
Who provides flserver.australian-cancer-data.network?
- Application / service owner: The FL server application is developed and operated by the ACDN research group (UNSW and Ingham Institute) as part of ACDN federated learning studies.
-
Infrastructure provider:
The VM behind
flserver.australian-cancer-data.networkruns on the Nectar Research Cloud, operated by the Australian Research Data Commons (ARDC) and its node partners (Australian universities). Nectar is a national research cloud used widely by Australian research organisations for secure research computing.
Where are the server and data located?
- FL server VM: hosted in Australia on the Nectar Research Cloud (within an Australian university data centre / Nectar node).
- Clinical and research datasets: stored only within participating hospital environments.
- Data on Nectar VM: no PHI and no de-identified clinical datasets are stored on the Nectar VM. Only model parameters/updates are exchanged.
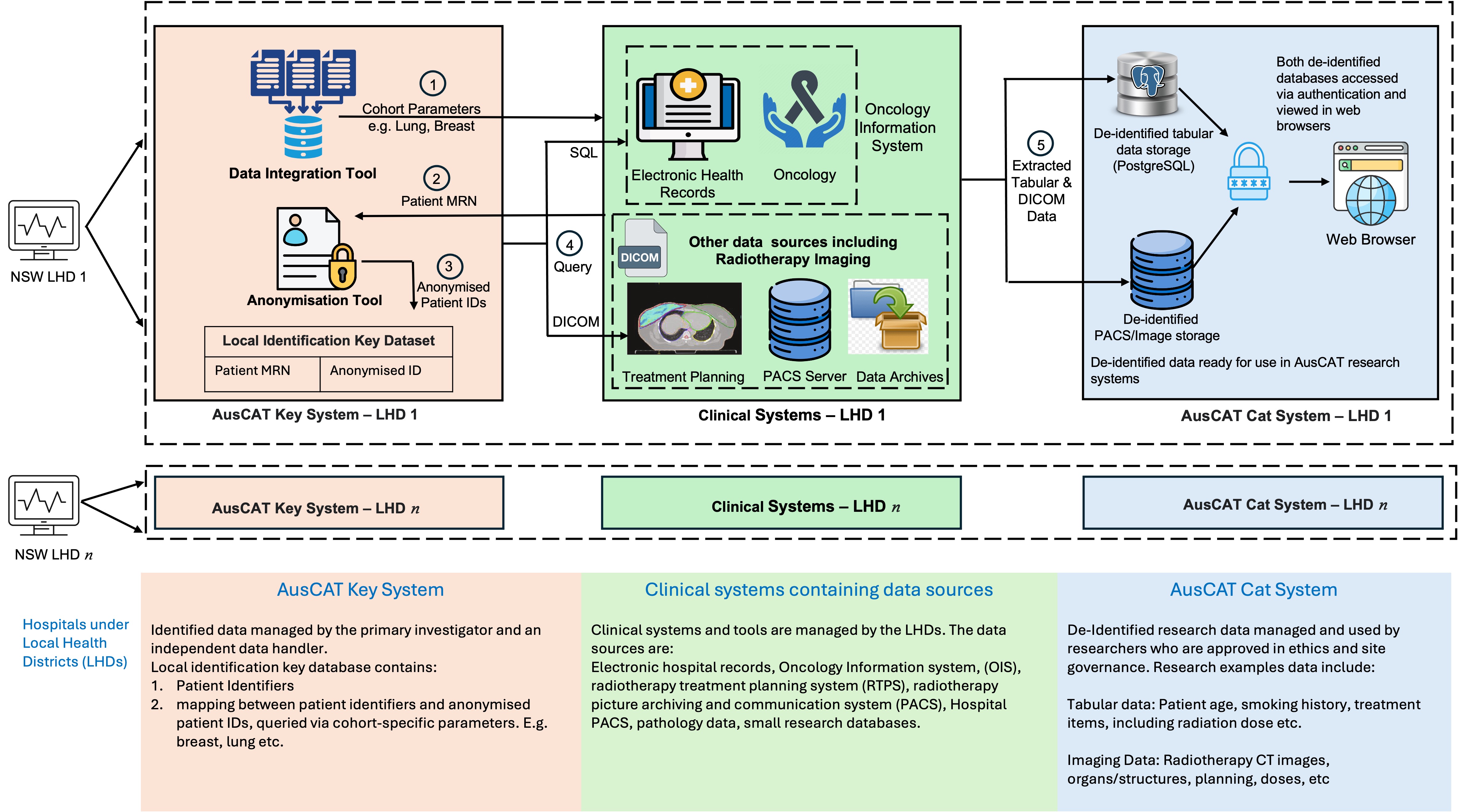
Data sources and architecture
Each participating hospital (under its Local Health District, LHD) identifies cohorts (e.g., breast or lung cancer) and performs local extraction from clinical systems.
- Clinical tabular data (OIS / EHR): extracted from systems such as MOSAIQ, ARIA, EPIC and associated electronic health records. Extraction is performed using SQL queries.
- Imaging and radiotherapy objects (TPS / PACS / archives): extracted from systems such as Pinnacle or Eclipse, PACS and clinical/research data archives. Extraction uses DICOM protocols and typically includes CT (and where relevant MRI/PET), RTSTRUCT, RTPLAN, and RTDOSE.
De-identification and separation of identifiable data
Identifiable patient information is managed within a dedicated local identification-key environment (separate VM/system). Access is restricted to authorised personnel according to local governance.
Identifiable data required for linkage is processed through an anonymisation pipeline using the RSNA MIRC Clinical Trials Processor (CTP) to produce de-identified datasets for research use.
ACDN hospital research environment
Following de-identification, research datasets are stored within a separate research VM/environment at each site:
- Tabular clinical data: stored in secure databases (e.g., PostgreSQL; managed via tools such as pgAdmin).
- Imaging data: stored in site PACS / image storage services (e.g., Orthanc DICOM server).
- Access controls: only authorised researchers/technical staff with governance approvals can access research datasets.
Researchers typically access services via controlled interfaces and secure ports with authentication enforced at multiple layers.
Federated Learning (FL)-based model communication
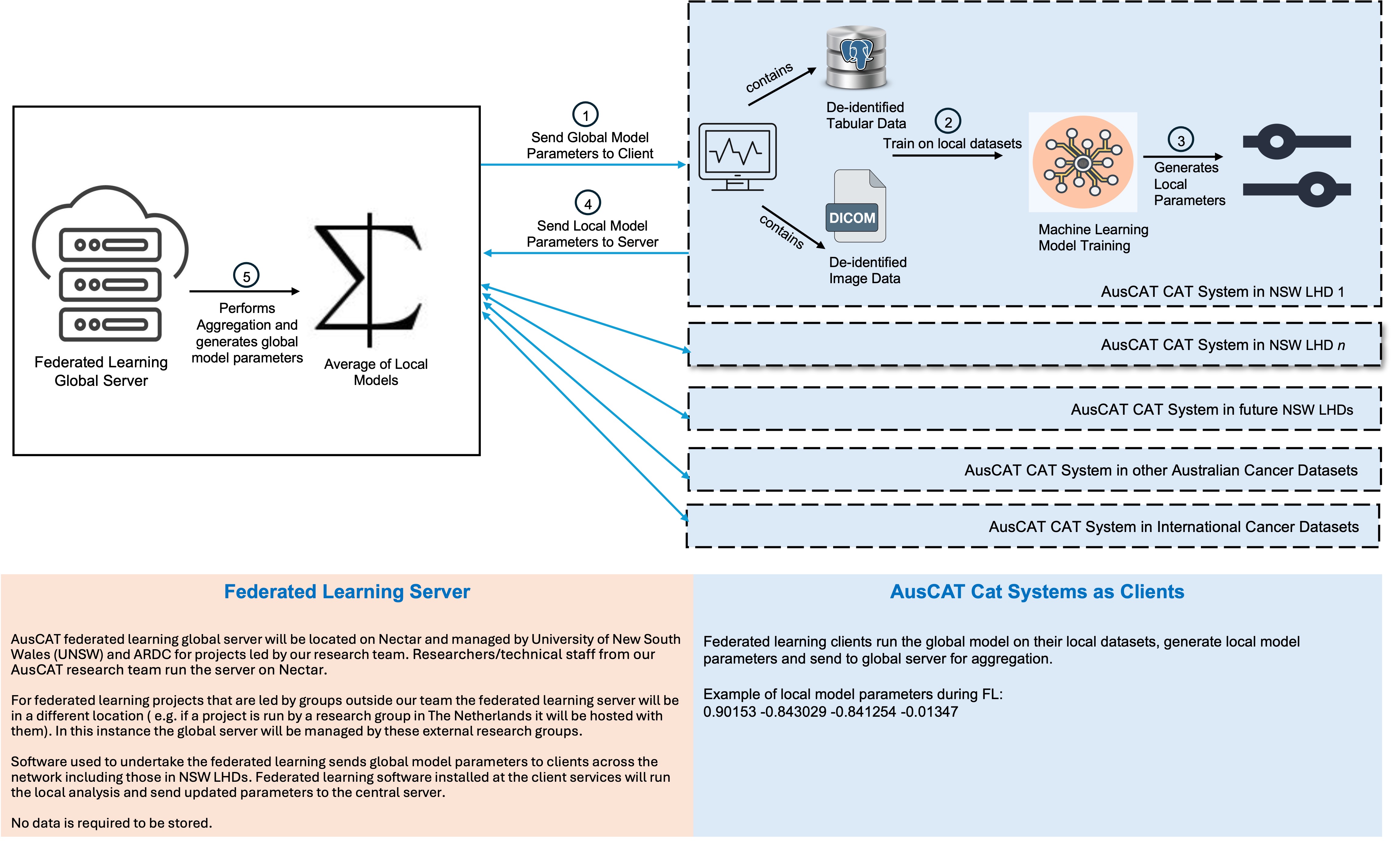
Once de-identified datasets are prepared and stored locally within each hospital, Federated Learning is used to collaboratively train models without transferring patient data outside the hospital network.
What is sent from each client to the FL server
- Model parameters / gradients: numeric arrays (floating-point values) representing the local model state after training.
- Aggregate metrics: summary statistics (e.g., loss/accuracy) used to monitor training.
What is NOT sent
- No DICOM images
- No clinical tables or exports
- No identifiers (names, MRNs, DOB, etc.)
No Patient Health Information (PHI) is transferred. Raw clinical datasets remain within each hospital. The FL server performs aggregation of model updates only.
Federated learning server and client interaction
Networking requirements (domain + port 5010)
FL client communication from participating hospital research VMs may be restricted by local proxy/firewall policies. For federated learning participation, sites may need to allow outbound connectivity to a destination domain and port.
- Destination domain:
flserver.australian-cancer-data.network(resolves to115.146.84.72) - Federated learning service port:
5010/tcpWhitelisting target - Web portal:
443/tcp(HTTPS) and80/tcp(HTTP → HTTPS redirect; also used for automated certificate renewal)
The requested whitelisting enables encrypted model-update exchange between clients and the server without transmitting PHI. Additional servers/domains may be used for externally run projects as needed, following the same principle: only model updates are exchanged.
Contacts
- Professor Lois Holloway — Lois.Holloway@health.nsw.gov.au
- Fahim Irfan Alam — fahim.alam1@unsw.edu.au
- Amir Anees — a.anees@unsw.edu.au
- Sasha Barisic — sasha.barisic@health.nsw.gov.au